Bacterial Cell Surfaces and Pathogenesis
Description of the different lines of research being followed by members of the group.
Brief description of the lines of research followed by members of the research team:
| The macromolecule of peptidoglycan (or its small components) seems to be able to induce an inflammatory response in different hosts. We believe that in order to understand how this macromolecule is sensed by the infected host we will need to learn how this molecule is synthesized, organized, modified and degraded during the bacterial cell cycle. There are four essential questions that represent the different research lines being followed in the Bacterial Cell Surfaces and Pathogenesis laboratory:
We use Staphylococcus aureus and Streptococcus pneumoniae, Gram-positive bacterial pathogens, as model organisms to study how the metabolism of the bacteria cell surface, mainly cell wall synthesis and turnover, can modulate a response of the innate immune system of the infected host. |
||
 |
||
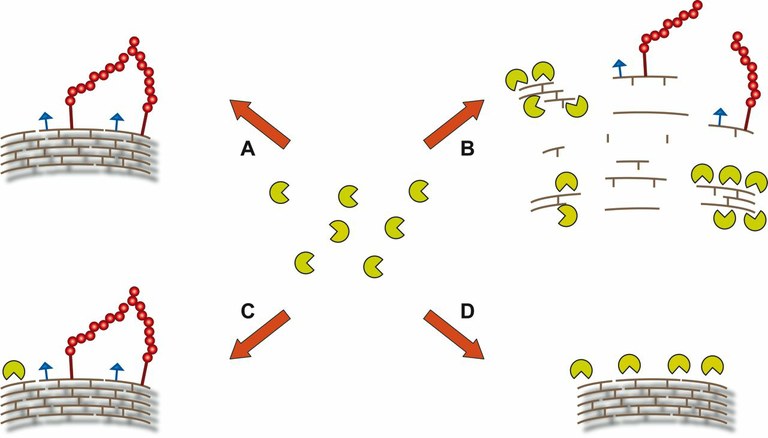
| Possible outcomes when a protein (yellow pacman) from the host encounters peptidoglycan in live bacteria: A) recognition is impaired by bacterial surface proteins or other polysaccharides; B) recognition occurs only when degradation of bacterial cell wall allows the release of peptidoglycan fragments that are recognized by the protein; C) recognition of peptidoglycan occurs only at specific sites of the bacterial surface or D) recognition is only possible with "naked" peptidoglycan, when no proteins or other polysaccharides are attached. |



