Abn2
Identifying the molecular basis that underlie the enzymatic mechanism of the arabinanase Abn2
Collaboration between the Laboratories of Isabel Sá-Nogueira, Isabel Bento and Cláudio M. Soares.
Financed project: PTDC/AGR-AAM/102345/2008 "Breaking Down The Wall - Microbial Hemicellulases for saccharification"
The plant cell wall is structurally complex and biologically recalcitrant. Microorganisms, in particular saprotrophs, play a fundamental role in the decomposition processes of plant biomass and secrete numerous polysaccharide-degrading enzymes that attack cellulose, hemicellulose, and pectin. Mobilization of plant biomass for chemical and fuel production is a major biotechnological challenge of the 21st century, and the application of polysaccharide-hydrolyzing enzymes in biomass saccharification is promising.
Endo-1,5-a-L-arabinanases are glycosyl hydrolases that are able to cleave the glycosidic bonds of the a-1,5-l-arabinan with the concomitant release of arabino-oligosaccharides and l-arabinose, one of the constituents of hemicellulose. Two different extracellular endo-1,5-a-L-arabinanases have been isolated from Bacillus subtilis BsArb43A and BsArb43B (AbnA and Abn2, respectively). Abn2 shows a low sequence identity with the other 1,5-a-L-arabinanases, is a much larger enzyme that comprises an N-terminal and C-terminal domain. The N-terminal domain is the catalytic domain and displays the characteristic beta-propeller fold common to these type of enzymes. We determined the structure of this enzyme (de Sanctis et. al. 2010) and found an extra C-terminal domain that comprises 103 amino acids not found in other arabinanases with known three-dimensional structure .
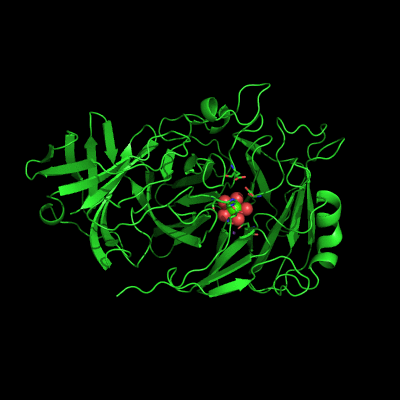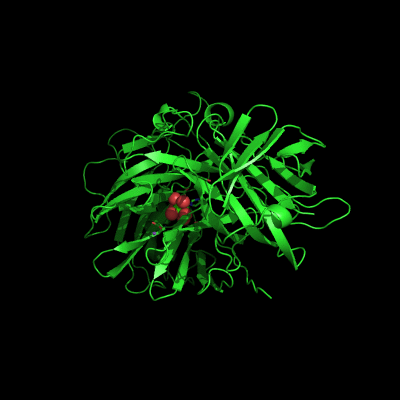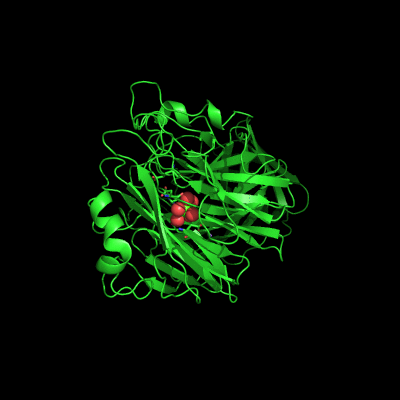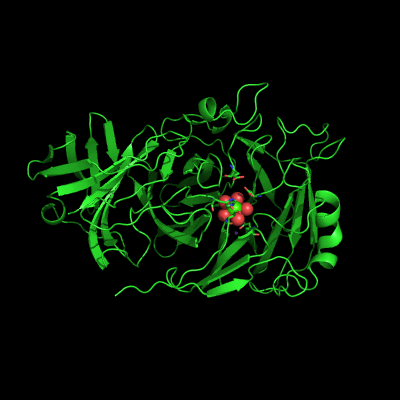
Abn2 from Bacillus subtilis (de Sanctis et. al. 2010)
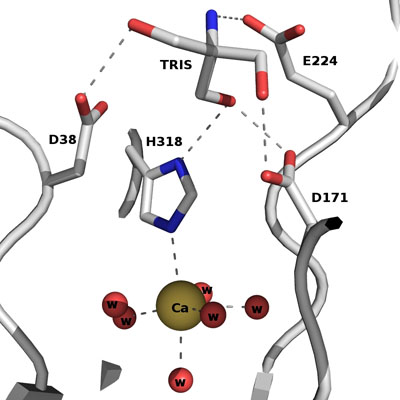
The active site of Abn2 is located in the deep cavity at the centre of the beta-propeller and comprises three carboxylate residues: Asp38, Asp171, and Glu224. These three acidic residues, which are conserved in all members of Glicoside Hydrolase families 32, 43, 62 and 68, are responsible for the general acid catalysis that leads to the hydrolysis of the glycosidic bond. Further down in the active cleft a calcium ion is observed, hepta-coordinated to six water molecules and a histidine ligand. The presence of this ion has been also reported in other structures of arabinanases but its role is yet to be determined.
ABN2 active site
In this research project we aim to understand the molecular mechanisms in Abn2 and to identify the key residues that are important for substrate binding and stabilization in the binding cleft. We are using a combination of Molecular Biology, Biochemistry, X-ray Crystallography and Biomolecular modelling techniques in order to achieve this goal.

MD simulation of ABN2 in water (0-100ns)
We are hiring. If you are interested in this project, please send us an email.
Recent relevant publications:
Daniele de Sanctis, Isabel Bento, Jose Manuel Inacio, Sonia Custodio, Isabel de Sa-Nogueira, Maria Armenia Carrondo (2008) Acta Cryst. F64, 636–638
Daniele De Sanctis, José M. Inácio,Peter F. Lindley, Isabel De Sá-Nogueira, Isabel Bento (2010) "New evidence for the role of calcium in the glycosidase reaction of GH43 arabinanases", FEBS J., 277, 4562–4574.