Looking critically at the landscape
Oeiras, 21.12.09
The Molecular Simulation Laboratory at ITQB has devised a new approach to characterize and understand the conformational behaviour of biomolecules. The work consists of a critical investigation of the energy landscape of a small peptide and has been selected to illustrate the cover of one of the December issues of the Journal of Physical Chemistry B.
Nowadays, structural changes of biomolecules are often addressed using the concept of energy landscapes. The idea is to think of your protein or peptide wandering around a mountain landscape in a way that reflects its structural preferences – valleys correspond to favoured low-energy conformations and peaks to high-energy disfavoured ones. Rather than just sliding downhill into valleys, the molecule is actually kicked around by thermal collisions, occasionally visiting mountain passes and peaks, and ends up populating the whole landscape with a pattern that reflects the energy of each region. The landscape concept provides an intuitive and appealing framework to rationalize protein folding and other structural changes in biomolecules. Therefore, much effort is being put into deriving energy landscapes from experimental and/or simulation data.
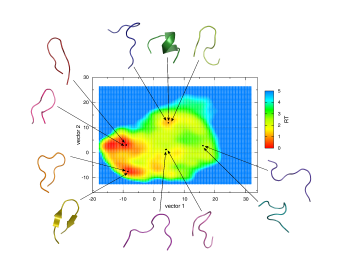
Most energy landscapes found in the literature are expressed as a function of two parameters, resembling the latitude/longitude of a real mountain landscape. Hence, it is possible to display energy landscapes as a familiar two-dimensional topographic map, with different energy levels indicated by contour lines or colours. This simple approach allows immediate recognition of the major groups of conformations (valleys) and the energy barriers (mountain passes) between them, but it implicitly assumes that all the conformations can be arranged meaningfully using just two parameters. The ITQB researchers found that this is actually impossible even for a small peptide.
|
|
Considering a ten-residue peptide (an arginylglutamate repeat motif found in several proteins and presumably involved in protein-protein interaction), ITQB researchers followed the usual two parameters procedure and found that the resulting topographic map is highly misleading and basically useless. Although a well-defined and smooth landscape was obtained, a closer inspection revealed several valleys with strikingly different peptide conformations. As it turns out, seven or eight parameters were needed to get a meaningful energy landscape that does not lump dissimilar conformations into the same region. Also, as researchers found, different methods of measuring conformation similarity are not equivalent and actually are suited for different purposes (for example, molecular recognition versus conformational kinetics). |
Overall, the present study shows that the usual two-dimensional energy landscapes should be regarded carefully. While surely appealing to rationalize your data, these approaches can also be deeply misleading. Fortunately, there are theoretical methods to check how meaningful they are.
Original article
The Journal of Physical Chemistry B 2009 113 (49), 15989-16001
Conformational Analysis in a Multidimensional Energy Landscape: Study of an Arginylglutamate Repeat
Sara R. R. Campos, António M. Baptista