Clockwork bacteria
Oeiras, 16.09.2016
Even in organisms as simple as bacteria, cell differentiation requires a precise regulation of gene expression. Researchers from the Microbial Developmental Lab and collaborators unveiled the mechanism for the activation of the late stages of sporulation in Clostridium difficile, which has revealed to be more complex than expected. The work was published yesterday in PLOS Genetics.
Clostridium difficile (or C. difficile) is the most important pathogen agent in hospitals, causing an intestinal syndrome whose symptoms can range from antibiotic-associated diarrhea to potentially deadly complications. Its ability to sporulate allows its dissemination and persistence in the host or hospital environment. Also, nowadays, community-associated or food chain-associated dissemination is becoming a major concern, and new therapeutics are urgently needed.
When C. difficile are exposed to adverse environmental conditions, they are able to differentiate into spores, a highly resistant type of cell. The process of spore formation involves two cell types – the forespore, which will give rise to the spore, and the mother cell, inside which the spore is formed. During this process more than 10% of the genome is recruited, which means that hundreds of genes have to be expressed in the right cell at the right time. To make sure spore formation is done properly nothing can be left to chance, regulation is key to understand how the process is done with precision and rigor.
The formation of bacterial spores is controlled by specific proteins – the sigma factors - that act as switches, activating and repressing dozens of genes. There are four different sigma factors (F, G, K, and E) with a very precise action in time and space. In this work, researchers from the Microbial Development Lab have focused on sigmaK a mother cell specific regulator, responsible for the assembly of the spore surface.
In 2015, researchers from Microbial Development Lab and collaborators already had discovered a mechanism by which the time of activation of a sigma factor is precisely controlled. The mechanism involved an anti-sigma factor, which functions to ensure that its target sigma factors remain inactive when and where necessary.
Now, a different type of mechanism was shown to control the timing of activation of sigmaK in the human pathogen Clostridium difficile. SigmaK is a mother cell specific regulator, that activate the final stages of spore morphogenesis and has a key role in formation of the spore surface layers. Proper formation of the spore surface layers is critical for host colonization by spores.
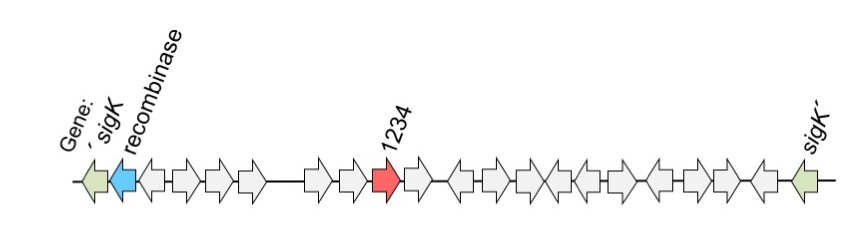
Figure 1: sigK gene (blue; coding for sigmaK) interrupted by a segment of DNA called skin (for sigmaK intervening sequence). The skin element contains the gene for a recombinase and the CD1234 protein, both of which are required for skin excision, and the generation of a functional sigK gene.
The gene coding for sigmaK is interrupted by an intervening sequence, called skin, carrying many genes. One of them codes for a recombinase which is required to excise the intervening sequence producing a functional sigmaK gene. Excision can only occur in the mother cell, so that the spore carries a complete genome to the next generation. However, in the work now published, researchers have realized that the recombinase is also produced in the spore. So the question now was: what restricts excision to the mother cell?
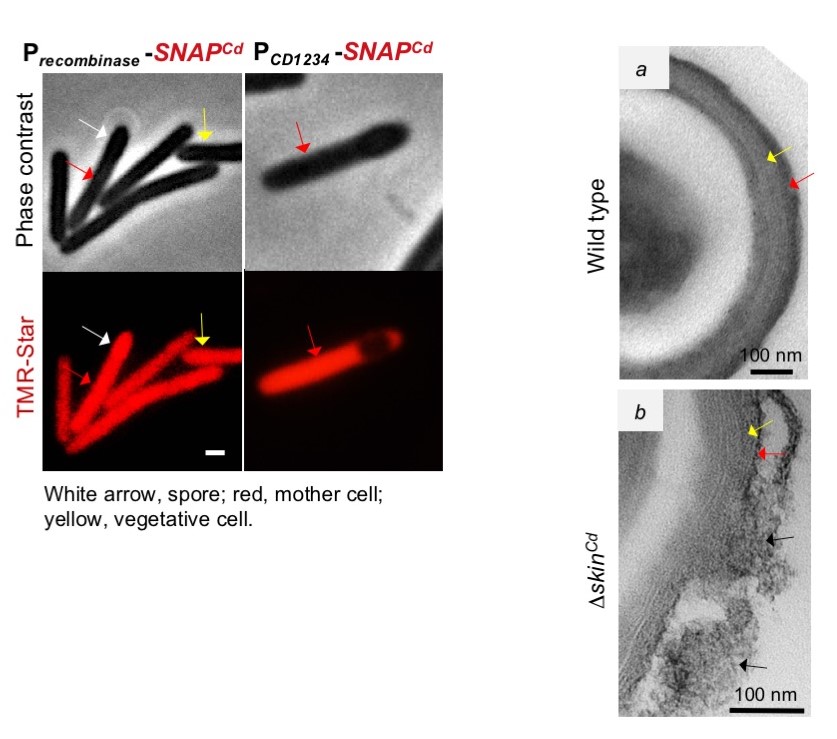
Figure 2 (left): Expression of the recombinase gene (red) occurs in vegetative cells, and in both the mother cell and spore of sporulating cells. Expression of CD1234, however, is confined to be mother cell.
Figure 3(right): the surface of wild type spores, and the disorganized surface of spore made by a strain from which skin was deleted, and that as the results show premature activity of sigmaK.
Researchers found that the expression of another skin gene, that they have called CD1234, is itself restricted to the mother cell. Once identified, researchers have found very interesting properties of this novel protein: CD1234 controls the recombinase enzyme. By itself, recombinase is able to introduce DNA into the genome, and even to invert an inserted sequence, but excision only occurs with the help of CD1234. The recombinase and CD1234 can be used as tools for genetic engineering in vivo and in vitro. Deletion of the skin sequence bypasses the need for the recombinase and for CD1234 in spore development. It also results, however, in premature activity of sigmaK, and in profound alterations of the spore surface, which most likely will influence the ability of spore to colonize their hosts. Thus, skin excision is a mechanism to ensure the correct time of sigmaK activation and proper spore morphogenesis. Since some naturally occurring epidemic strains lack a skin element, the authors speculate that these strains have manipulated the timing of sigmaK activation to generate variation and exploit new ways of infecting their hosts.
ORIGINAL ARTICLE
PLoS Genet 12(9): e1006312. doi:10.1371/journal.pgen.1006312
Mónica Serrano, Nicolas Kint, Fátima C. Pereira, Laure Saujet, Pierre Boudry, Bruno Dupuy, Adriano O. Henriques, Isabelle Martin-Verstraete
THE LAB AND THE RESEARCHERS

Adriano Henriques (back row, on the right) coordinates the Microbial Development Lab. The lab has been interested in understanding the mechanisms of bacterial spore formation, since they have a central role in persistence of sporeforming bacteria, including pathogens such as C. difficile, in the environment, infection, recurrence and transmission of diseases. The group has been focusing on the assembly of the spore surface layers.
Mónica Serrano (front row, second on the right) is FCT Investigator since 2014, and has been working on the Microbial Development Lab. She has recently been awarded 2,5 million euros to co-coordinate the project ONEIDA, to prevent and control bacterial infections in Portuguese hospitals.
On Social Media
Facebook, Twitter and Linkedin
In the news
- As bactérias são mais precisas que relógios, Sul Informação, 16.09.2016
- Bactérias são mais precisas que relógios, Elvas News, 16.09.2016
- Bactérias são mais precisas que relógios, Notícias do Nordeste, 17.09.2016
-
Bactérias são mais precisas que relógios, Penacova Actual, 19.09.2016
-
Bactérias são mais precisas que relógios, Jornal Aurinegra, 19.09.2016







